| Topic | Key Points |
|---|---|
| Title | iDASH 2025 Track 1 – Secure Evaluation of DNA-binding Classification CNN |
| Objective | Securely evaluate a provided CNN on encrypted DNA sequences using homomorphic encryption (HE); return encrypted prediction identical to plaintext run. |
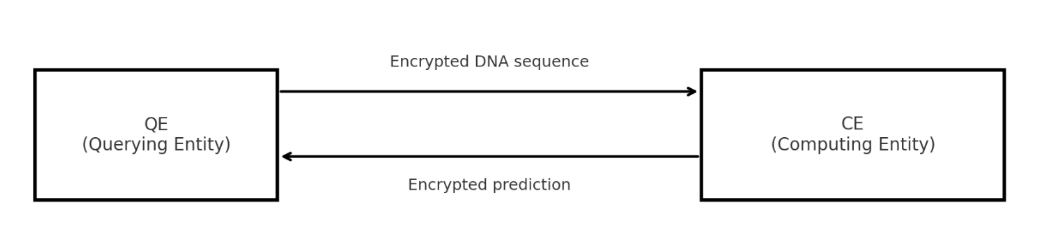
| Parties | QE (querying entity – owns sensitive DNA) & CE (computing entity – owns model). |
| Model | Torch CNN with conv + pooling + FC layers; binary protein-DNA binding output; no retraining needed. |
| Input | 2-column, tab-delimited file; each sequence: 200 nt (A,C,G,T). |
| Output | Participant-defined text file; one score/vector per sequence (after QE decryption). |
| Provided | • Torch model + docs • Example file (1 000 seqs/classes) for format only. |
| Technical Rules | • Implement/approximate every layer (incl. final) under HE Std (Albrecht-19). • Only one linear scaling allowed before/after decryption. • No explicit QE/CE traffic simulation needed. |
| Evaluation | Test set: 2 000 sequences. Metric: auROC / exp(wall-time / 20 min); >60 min wall-time ⇒ disqualified. |
| Deliverables | Code/binaries + documentation; BSD 3-Clause license; submission link (TBA). |
| Timeframe & Limits | ≤60 min wall-time for full evaluation; faster yields better score via exponential penalty. |
| Data/Code Usage | No dataset sharing; no publications before workshop; abide by shared licenses. |
| Support/FAQ | FAQ Document |
| Docs & Data link | Documentation and Data |
Track 2: Access Request Recording and Querying for Biomedical Datasets
Goal
To develop blockchain-based smart contracts for managing biomedical data requests, to facilitate the continuity/efficiency of biomedical, clinical, and genomic research [1].
Challenge
Multiple smart contracts must be implemented to manage both data (i.e., data requests) and dictionaries (e.g., list of principal investigators).
All data/dictionaries and intermediary data/dictionaries (e.g., index or cache of the original data/dictionaries) must be stored entirely on-chain via smart contracts (i.e., no off-chain data storage is allowed).
We will provide the skeleton of the smart contracts.
The system must manage data requests from research institutions while enforcing specific data access requirements around dataset ownership, Principal Investigator (PI) credentials, and Data Use Agreements (DUA).
Each participant can determine how each insertion is represented and stored in the smart contracts.
Participants can implement any algorithm to store, retrieve and present the data/dictionaries correctly and efficiently.
Users should be able to query the data from any of the blockchain nodes.
Evaluation Criteria
The data access request system will need to demonstrate satisfactory performance (i.e., 100% accurate query results) on a test dataset, which will be different from the training set provided online.
We will evaluate the efficiency of each solution using insertion and query times.
We will insert data requests for a specified time frame before conducting the queries.
We will only grade the accuracy and storage/retrieval speed on data but not dictionaries (i.e., we will pre-store the dictionaries on-chain before evaluating participants’ smart contracts).
Experimental Setting
Given a set of institutional biomedical data requests, design a time/space efficient data structure and mechanisms to manage (i.e., store and retrieve) these requests based on Ethereum Solidity smart contracts.
Additional Rules
The submission should include 3 files of Solidity source code per team.
All 3 smart contracts must be implemented.
Reusing any external code/library must follow the license agreement for both the code/library and our track (please see below License section for more details), and the reusing code blocks must be clearly and explicitly cited using Solidity comments.
One person can only participate in one team. All team members' names and emails must be listed as in Solidity comment in the submission code file and team membership cannot be changed after the submission due date. Although it is allowed to submit multiple times before the due date, only the last submission will be evaluated. If there are solution-wise communications cross teams, it must also be disclosed in comments. The solutions of the teams not following the rules above will not be evaluated for fairness consideration.
Data Usage and Publication Agreement
By registering and/or participating in this challenge and receiving restricted access to the challenge dataset, members of all teams agree to abide by the following rules of data usage: (1) They will not share the challenge dataset with others. (2) They will not use the challenge dataset in any publications until after the iDASH 2025 Workshop concludes. These are set up to ensure fairness among the participating teams.
License
This track requires every participating team to share their code and/or binaries under the BSD 3-Clause License Open Source license. The track co-organizers will upload all submitted code and/or binaries to a GitHub repository under the BSD 3-Clause License right after the competition results are announced. By submitting the code and/or binaries, the participants automatically consent to allow the track co-organizers to release the code and/or binaries under the BSD 3-Clause License.
Data Skeleton
FAQ:
For questions and clarifications, please check and post on our FAQ here
References
- Yu Y, Edelson M, Pham A, Pekar JE, Johnson B, Post K, Kuo T-T. Distributed, immutable, and transparent biomedical limited data set request management on multi-capacity network. Journal of the American Medical Informatics Association. 2024. doi: 10.1093/jamia/ocae288. PubMed PMID: 39569448.